Our Research Vision
To expand precision medicine from the current gene-centric process to one that considers the complexity and dynamics of proteins. We strive to develop new diagnostic and therapeutic approaches based on the altered proteins unique to cancer.
Precision Oncology
Personalized Oncology & Molecular Pathology
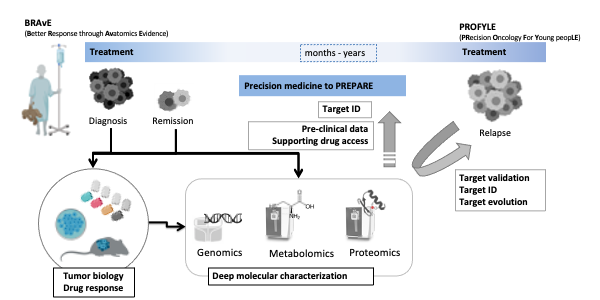
Finding kinder, more targeted treatments is a key priority in childhood cancer research. To advance precision oncology we co-founded the BRAvE program at BCCH and co-lead the proteomics node in the national PROFYLE initiative.
- Establishing real-time proteomic profiling and Molecular Tumour Board reporting.
- Studying drug response in patient-derived cells, CAM, and mouse models.
- Defining ‘normal’ protein ranges in children and young adults (CAYA).
- Developing biomarker panels and pediatric cancer-specific MS assays.
Immunotherapy
Cell Surface Proteomics & New Targets
Pediatric malignancies have a far lower mutation rate than adult cancers, requiring a different treatment approach. We have identified a new class of neoantigens that shows great potential in pediatric cancers.
- Combining cell-surface proteomics of cancer models and patient biopsies.
- Evaluating neoantigen presence throughout disease progression.
- Developing lead compounds to specifically target these neoantigens.
Microenvironment
Cell-Cell Communication
Cells secrete bioactive proteins that trigger responses in neighboring cells. We integrate cell biology, biochemistry, and proteomics to study extracellular signaling networks.
- Established the chicken chorioallantoic membrane (CAM) model for accessible mechanistic studies.
- Investigating how secreted proteases (Cathepsins and MMPs) remodel paracrine signaling.
Data Integration
Proteoform Function Deconvolution
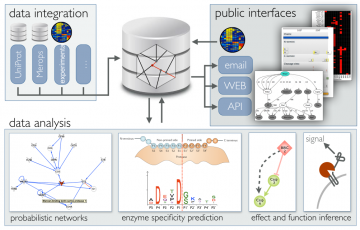
To harness the full potential of proteoforms, we need tools to define the landscape and confirm that proteoform-level measurements exceed protein-level measurements.
- Developing algorithms for multi-scale data integration and pattern recognition.
- Building new biological knowledgebases to guide personalized treatment decisions.
Mass Spectrometry
New Technologies & Methods
Advances in clinical specimen analysis are enabled by breakthroughs in sample processing and protein biochemistry.
- Improving existing and developing new wet-lab and computational technologies.
- Enabling comprehensive investigation of small samples and complex systems.