
This word cloud represents the key terms from our lab’s publications as indexed by Google Scholar (up to October 2025). It was generated using this tool.
How to Read This Page: Lab members are underlined in the author lists, and the Principal Investigator is marked in bold.
Preprints – Feedback welcome!
ProteoForge: An Imputation-Aware Framework for Differential Proteoform Discovery in Bottom-Up Proteomics
bioRxiv, 2025 (under review at Journal of Proteome Research)
More pre-prints on the way, please check back soon.
Peer-reviewed publications

Proteomics and personalized PDX models identify treatment for a progressive malignancy within an actionable timeframe
EMBO Molecular Medicine, 2025

Statistical Testing for Protein Equivalence Identifies Core Functional Modules Conserved across 360 Cancer Cell Lines and Presents a General Approach to Investigating Biological Systems
Journal of Proteomics Research, 2024
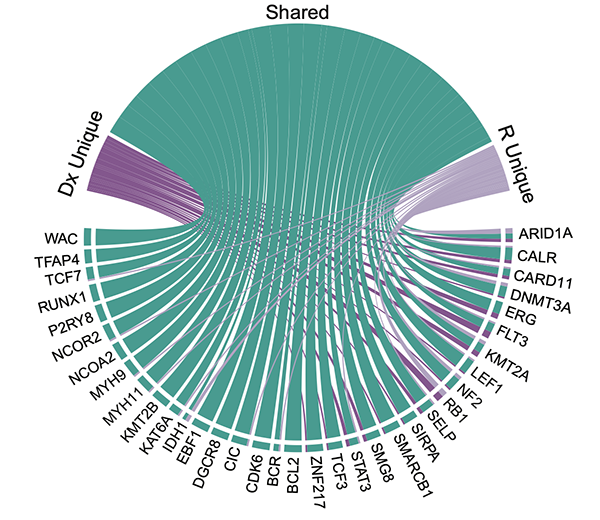
Targetable lesions and proteomes predict therapy sensitivity through disease evolution in pediatric acute lymphoblastic leukemia
Nature Communications, 2023
Pathogenic BRCA1 variants disrupt PLK1-regulation of mitotic spindle orientation
Nature Communications, 2022
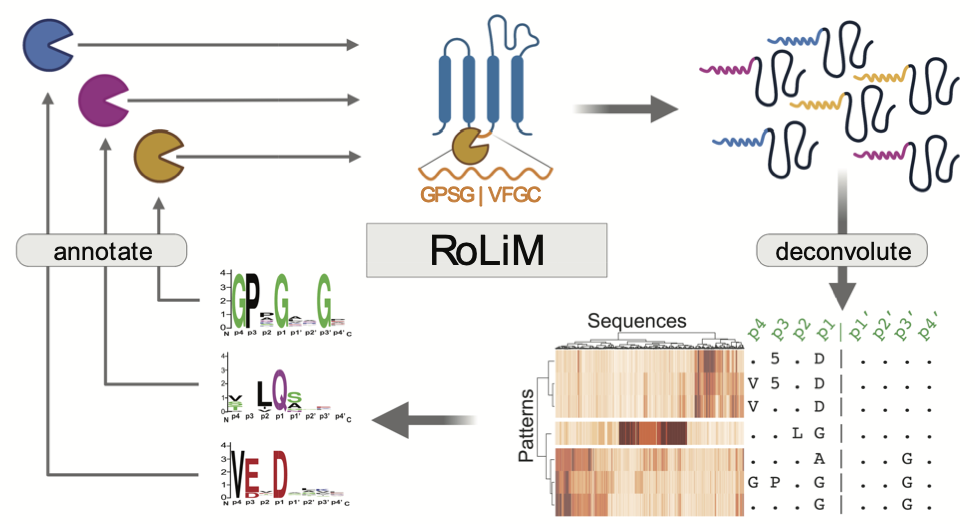
Sensitive identification of known and unknown protease activities by unsupervised linear motif deconvolution
Analytical Chemistry, 2022
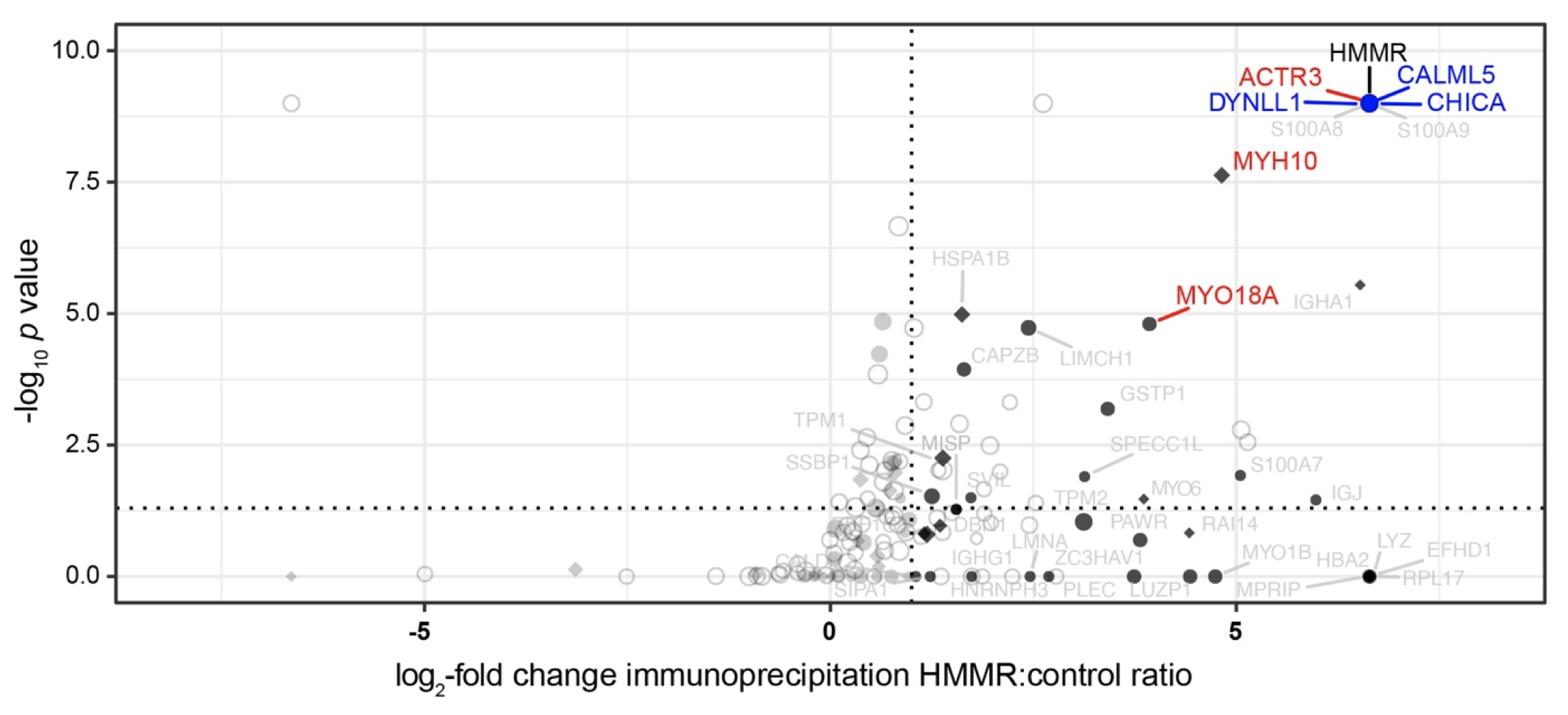
Modification of BRCA1-associated breast cancer risk by HMMR overexpression
Nature Communications, 2021
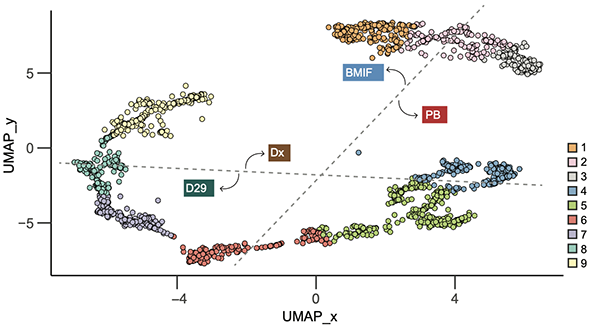
Multi-omic profiling of the leukemic microenvironment shows bone marrow interstitial fluid is distinct from peripheral blood plasma
Exp Hematol Oncol, 2022
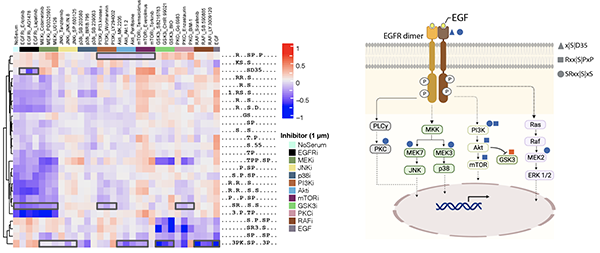
Robust unsupervised deconvolution of linear motifs characterizes 68 protein modifications at proteome scale
Scientific Reports, 2021
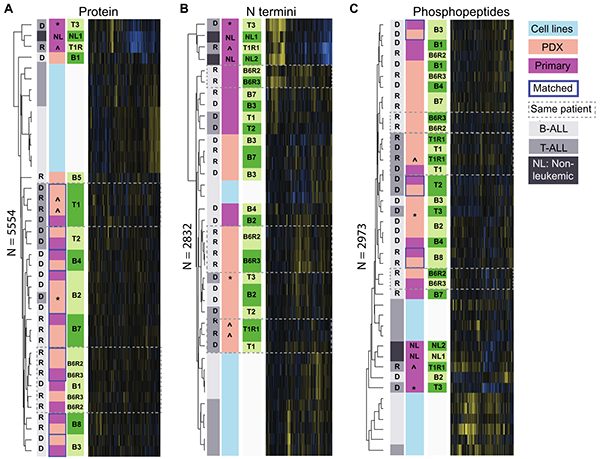
PDX models reflect the proteome landscape of pediatric acute lymphoblastic leukemia but divert in select pathways
Journal of Experimental and Clinical Cancer Research, 2021

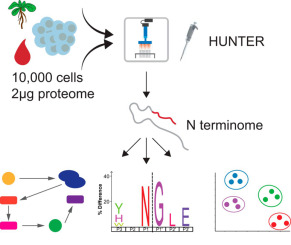
Sensitive determination of proteolytic proteoforms in limited microscale proteome samples
Molecular & Cellular Proteomics, 2020
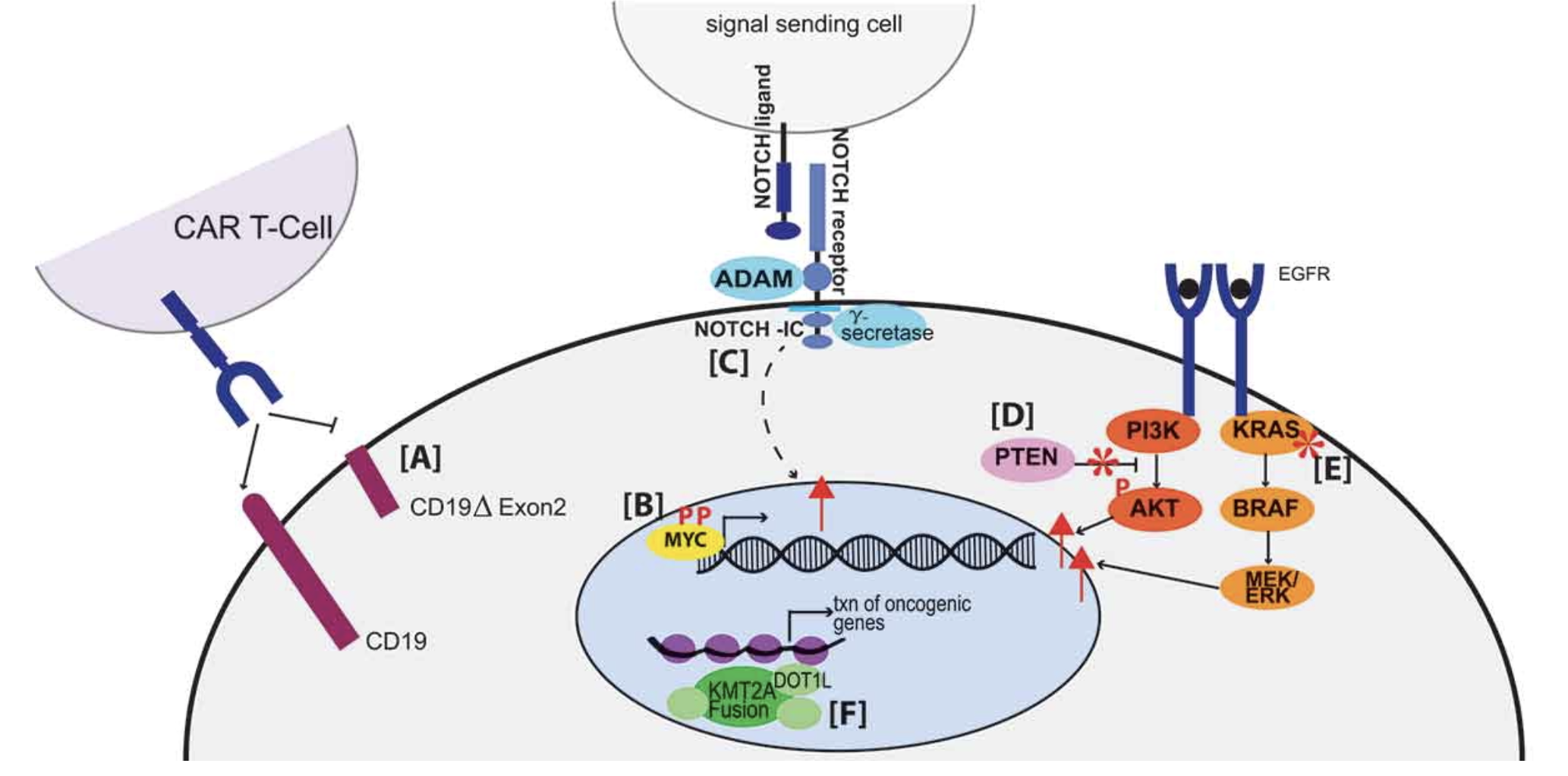
Origins and clinical relevance of proteoforms in pediatric malignancies
Expert review of proteomics, 2019
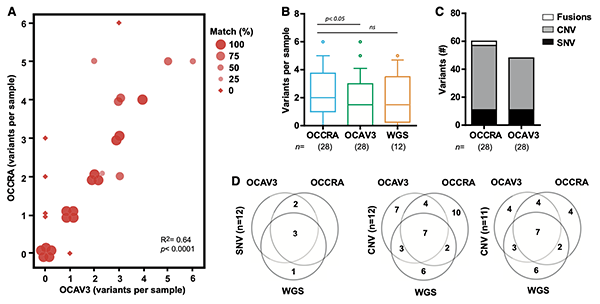
Tumor variant identification that accounts for the unique molecular landscape of pediatric malignancies
J National Cancer Institute (JNCI) – Cancer Spectrum, 2019
Other Manuscripts
- Yu H, Xing S, Nierves L, Lange PF, Huan T. Implementing quality control-based signal calibration in mass spectrometry-based metabolomics to overcome the error of fold change calculation caused by nonlinear ESI responses. Analytical Chemistry (2020)
- Richmond AP, Kloet F, Vaz FM, Kampen AHC, Uzozie A, Lange PF, Lin D, Kobor M, Graham E, Mostafavi S, Moreland P, Wasserman W, Karnebeek C, Kemp S, Engelen M. Multi-omic approach to identify phenotypic modifiers underlying cerebral demyelination in X-linked adrenoleukodystrophy. Frontiers in Cell and Developmental Biology
- Connell M, Chen H, Jiang J, Kuan C-W, Fotovati A, Chu T, He Z, Lengyell TC, Li H, Kroll T, Li AM, Goldowitz D, Frappart L, Ploubidou A, Patel M, Pilarski LM, Simpson EM, Lange PF, Allan DW, Maxwell CA. HMMR acts in the PLK1-dependent spindle positioning pathway and supports neural development. eLIFE (2017)
- Prudova A, Gocheva V, Auf dem Keller U, Eckhard U, Olson OC, Akkari L, Butler GS, Fortelny N, Lange PF, Mark JC, Joyce JA, Overall CM. TAILS N-Terminomics and Proteomics Show Protein Degradation Dominates over Proteolytic Processing by Cathepsins in Pancreatic Tumors. Cell Reports (2016)
- Fotelny N, Yang S, Pavlidis P, Lange PF#, Overall CM#. Proteome TopFIND3.0 with TopFINDer and PathFINDer: database analysis tools for the association of protein termini to pre- and post-translational events. Nucleic Acids Research (2015)
- Pitter F Huesgen, Philipp F Lange, Lindsay D Rogers, Nestor Solis, Ulrich Eckhard, Oded Kleifeld, Theodoros Goulas, F Xavier Gomis-Rüth & Christopher M Overall. LysargiNase mirrors trypsin for protein C-terminal and methylation-site identification. Nature Methods (2015)
- Nikolaus Fortelny, Jennifer Cox, Reinhild Kappfelhoff, Amanda E. Starr, Philipp F. Lange, Paul Pavlidis, Christopher M. Overall. Network Analyses Reveal Pervasive Functional Regulation Between Proteases in the Human Protease Web. PLoS Biology (2014)
- Philipp F Lange*, Pitter F Huesgen*, Karen Nguyen, Christopher M Overall. Annotating N termini for the Human Proteome Project: N termini differentiate stable processed protein species from degradation remnants in the human erythrocyte proteome. Journal of Proteome Research (2014)
- Huesgen PF*, Lange PF*, Overall CM. Ensembles of protein termini are sensitive and specific proteolytic signature biomarkers of disease. Proteomics – Clinical Applications (2014)
- Lange PF, Overall CM. Protein TAILS: When termini tell tales of proteolysis and function. Current Opinion in Chemical Biology (2013)
- Philipp F. Lange, Pitter F. Huesgen, Christopher M. Overall. TopFIND 2.0—linking protein termini with proteolytic processing and modifications altering protein function. Nucleic Acids Research (2012)
- Philipp F. Lange & Christopher M. Overall. TopFIND, a knowledgebase linking protein termini with function. Nature Methods (2011)
- auf dem Keller U, Bellac CL, Li Y, Lou Y, Lange PF, Ting R, Harwig C, Kappelhoff R, Dedhar S, Adam MJ, Ruth TJ, Bénard F, Perrin DM, Overall CM. Novel matrix metalloproteinase inhibitor [18F]marimastat-aryltrifluoroborate as a probe for in vivo positron emission tomography imaging in cancer. Cancer Research (2010)
- Philipp F. Lange, Lena Wartosch, Thomas J. Jentsch, Jens C. Fuhrmann. ClC–7 requires Ostm1 as a β–subunit to support bone resorption and lysosomal function. Nature (2006)
